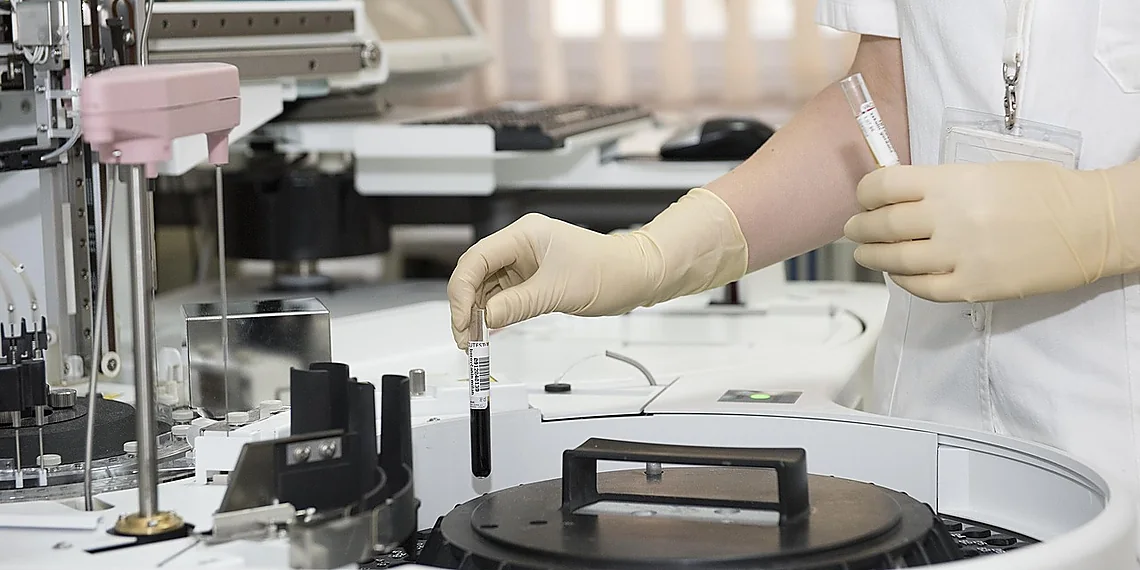
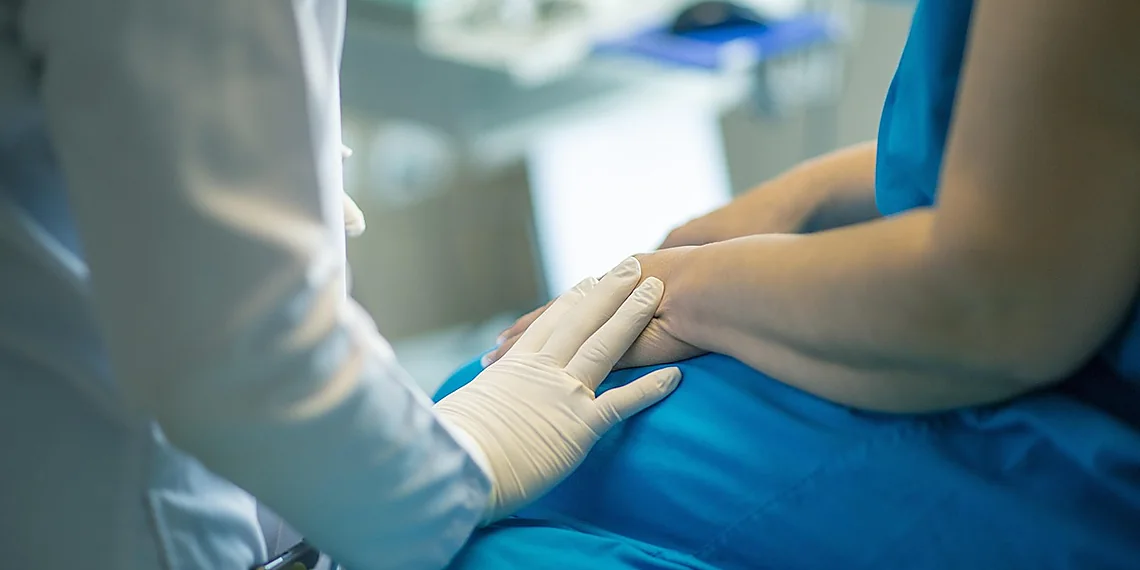
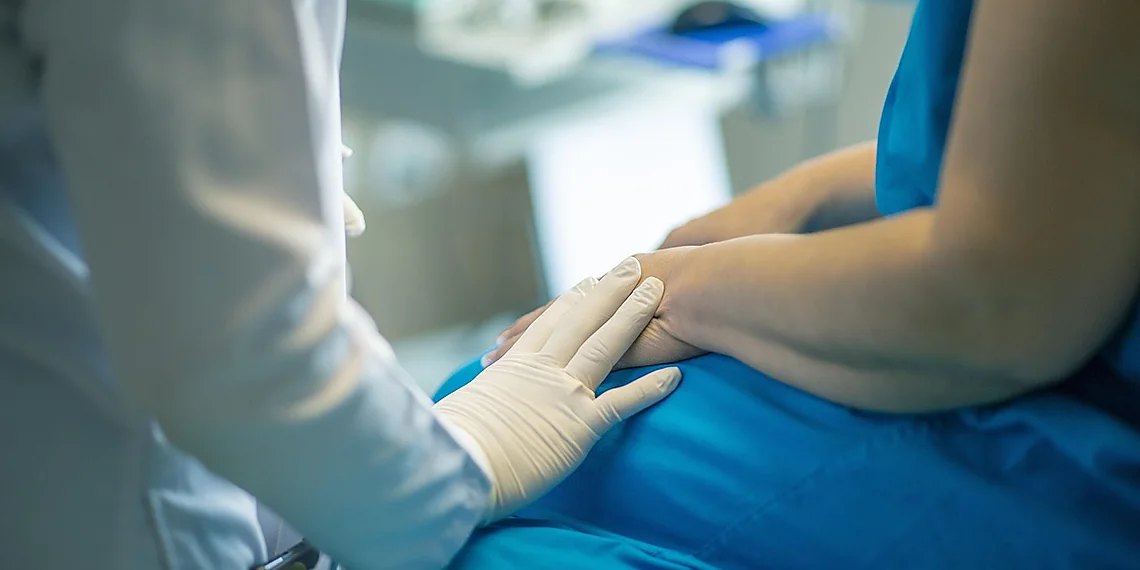
The relentless pursuit of breakthroughs in healthcare relies heavily on robust datasets, but accessing real patient information presents a formidable hurdle – privacy. Traditional medical research often faces significant delays and restrictions due to stringent regulations designed to protect sensitive personal health records, slowing the pace of innovation across numerous fields like drug discovery and personalized medicine. We’re entering an era where artificial intelligence offers a compelling solution to this longstanding problem: the ability to generate realistic, yet entirely fabricated, patient data for research purposes. This opens doors previously locked by ethical and legal limitations.
Imagine having access to thousands – or even millions – of detailed medical histories without compromising anyone’s confidentiality; that’s the promise of synthetic patient data. These aren’t simple simulations; they are complex digital representations mimicking real-world patient characteristics, including demographics, diagnoses, treatments, and lab results. Projects like Synthea have demonstrated impressive capabilities in generating these profiles, but crafting the underlying rules and algorithms to ensure accuracy and utility remains a significant challenge for researchers and developers.
The implications of this technology extend far beyond simply accelerating research timelines. Synthetic data has the potential to democratize access to valuable insights, enabling smaller institutions and startups to participate in vital studies previously unavailable to them. As AI continues to evolve, expect to see increasingly sophisticated methods for generating synthetic patient data, further blurring the lines between reality and simulation while upholding crucial privacy safeguards.
The Promise of Synthetic Patient Data
The healthcare industry is on the cusp of a significant shift, fueled by advancements in artificial intelligence – and much of that revolution hinges on something called synthetic patient data. Traditionally, accessing real-world medical records for research has been an arduous process, often stymied by stringent privacy regulations like HIPAA in the US and GDPR in Europe. These rules are vital to protect individuals’ sensitive health information, but they also create a bottleneck, severely limiting the ability of researchers to develop new treatments, diagnostic tools, and predictive models that could benefit countless people.
Synthetic patient data offers a compelling solution to this dilemma. Unlike real medical records which contain identifiable information, synthetic datasets are artificially generated to mimic the statistical properties of actual patient populations. These datasets can include everything from demographics and diagnoses to lab results and treatment histories – all without revealing any personal details. Because they don’t represent real individuals, synthetic data often falls outside the scope of strict privacy regulations, dramatically easing access for researchers and accelerating innovation.
The rise in popularity of tools like Synthea, a rule-based data generator, exemplifies this growing trend. Synthea creates realistic patient profiles based on predefined rules governing disease probabilities and other factors – allowing for complex scenarios to be simulated without compromising patient confidentiality. While creating these rules initially requires significant expertise and sample data, ongoing AI advancements are aiming to automate and refine this process, making synthetic data generation even more accessible and robust. This represents a pivotal moment in healthcare research where the promise of progress is increasingly tied to the responsible use of artificial datasets.
Ultimately, synthetic patient data isn’t about replacing real-world clinical trials or diminishing the importance of privacy; it’s about creating an enabling environment for innovation. By providing researchers with safe and readily available datasets, we can unlock new insights into disease progression, personalize treatment plans, and ultimately improve health outcomes – all while upholding the ethical principles that protect patient confidentiality.
Why Real Medical Data Is So Protected (and Hard to Access)

Accessing real patient medical data for research purposes is notoriously difficult due to stringent regulations designed to protect individual privacy. In the United States, the Health Insurance Portability and Accountability Act (HIPAA) establishes national standards for handling protected health information (PHI). HIPAA dictates how healthcare organizations must use and disclose patient data, requiring explicit consent or adherence to specific de-identification protocols that can significantly limit its utility for research.
Beyond HIPAA, similar privacy laws exist globally, such as the General Data Protection Regulation (GDPR) in Europe. These regulations impose significant hurdles for researchers seeking access to medical records, often necessitating lengthy approval processes from Institutional Review Boards (IRBs), data use agreements, and sometimes even requiring anonymization techniques that can compromise the quality and usefulness of the data for analysis. The complexity and cost associated with navigating these legal frameworks are substantial barriers.
The challenges in obtaining real patient data stem not only from regulatory compliance but also from ethical considerations regarding patient autonomy and confidentiality. Even when de-identified, there remains a risk of re-identification through linkage attacks or inference based on seemingly innocuous combinations of attributes. This inherent risk underscores the need for alternative solutions that can provide realistic data without compromising individual privacy – a key driver behind the growing interest in synthetic patient data.
Synthea: The Rule-Based Data Generator
Synthea has emerged as a prominent solution in the burgeoning field of synthetic patient data generation, offering a compelling approach to addressing privacy concerns while enabling valuable research and development opportunities within healthcare. Unlike methods relying solely on statistical mimicry, Synthea distinguishes itself through its rule-based architecture. This means that instead of simply replicating patterns observed in real datasets, Synthea operates by defining explicit ‘rules’ governing the entire lifecycle of a synthetic patient – from birth to death – ensuring a degree of coherence and realism often lacking in simpler generation techniques.
At the heart of Synthea’s functionality lies its sophisticated rule system. These rules dictate the probability of various events occurring throughout a patient’s simulated lifetime, factoring in elements like age, gender, family history (which itself is generated), and other relevant characteristics. For example, a rule might state that the likelihood of developing Type 2 diabetes increases with age and BMI. These probabilities aren’t arbitrary; they are designed to reflect known medical realities, although their precise values can be adjusted for specific research needs or to explore hypothetical scenarios. The result is a synthetic patient record complete with demographics, diagnoses, procedures, medications, lab results, and even social history – all generated in accordance with these predefined rules.
The beauty of Synthea’s rule-based approach lies not only in its ability to produce plausible data but also in its inherent privacy advantages. Because the system generates entirely new data points based on statistical principles rather than copying existing records, it inherently avoids direct identifiers and minimizes the risk of re-identification. While crafting these rules initially requires significant expertise – a deep understanding of medical conditions and their progression is essential – the resulting dataset can be used for tasks ranging from training machine learning models to testing clinical decision support systems without compromising patient privacy.
While Synthea represents a considerable advancement, it’s important to acknowledge the complexities involved in its implementation. Defining realistic and comprehensive rules requires substantial domain knowledge and often iterative refinement based on initial data outputs. Researchers are actively exploring methods to automate this rule creation process, as highlighted in recent work (arXiv:2512.14721v1), aiming to democratize access to synthetic patient data generation and further enhance the utility of platforms like Synthea for a wider range of healthcare applications.
How Synthea Works – Rules, Probabilities, and Patient Lifecycles

Synthea operates on a foundation of rules that dictate the progression and characteristics of each simulated patient’s lifecycle. These aren’t simple ‘if/then’ statements; instead, they are probabilistic assertions defining the likelihood of events occurring based on factors like age, gender, ethnicity, and existing conditions. For example, a rule might state that a 50-year-old male has a 15% chance of being diagnosed with hypertension within the next five years, while a similar rule for a 30-year-old female would have a lower probability.
The rules cover a wide range of patient attributes, from demographic information and medical history to lab results and medication prescriptions. Disease progression is also governed by these probabilistic rules; a diagnosis of diabetes might trigger subsequent rules outlining the likelihood of developing complications like neuropathy or retinopathy over time. This layered approach ensures that the synthetic data reflects realistic clinical scenarios and temporal dependencies.
Developing these rule sets requires significant domain expertise to ensure accuracy and realism. While Synthea’s initial versions relied heavily on manual definition, ongoing research focuses on automating this process using machine learning techniques trained on real-world patient datasets. The goal is to create a system where complex medical knowledge can be translated into effective rules for synthetic data generation with reduced human intervention.
Automating Rule Creation: A New Approach
Traditional synthetic patient data generation, particularly using tools like Synthea, relies heavily on meticulously crafted rules that dictate a patient’s journey – everything from age-dependent disease probabilities to medication adherence. These rules are the backbone of realistic synthetic datasets, allowing researchers and developers access to valuable medical information without compromising real patient privacy. However, creating these rules has historically been a significant bottleneck: it demands deep domain expertise and often requires painstaking manual effort to translate clinical understanding into precise, functional rule sets. This reliance on experts limits scalability and can introduce bias based on the expert’s perspective.
The research detailed in arXiv:2512.14721v1 introduces a groundbreaking innovation – automating this very process. Instead of relying solely on human experts, the new approach focuses on extracting statistical information directly from real-world data sources, such as cancer reports. This allows for the automatic generation of Synthea rules, effectively bypassing the need for extensive manual rule creation and reducing dependency on specialized knowledge. The core concept is to analyze observed patterns within existing datasets – for example, how the prevalence of a particular condition changes with age or other factors – and translate those statistical observations into corresponding Synthea rules.
The methodology involves a specific process where cancer reports are analyzed to identify key relationships and probabilities. For instance, if reports consistently show a correlation between a certain genetic marker and the development of a specific type of cancer at a particular age range, that information is then codified as a rule within Synthea’s framework. This doesn’t involve directly copying patient data; instead, it extracts the underlying statistical trends and translates them into probabilistic statements that guide the synthetic data generation process. The technical implementation focuses on identifying statistically significant patterns and converting them into a format compatible with Synthea’s rule structure, ensuring both accuracy and usability.
Ultimately, this automated approach promises to democratize access to synthetic patient data. By removing the expert dependency, it opens opportunities for broader participation in research and development, accelerates the creation of tailored datasets for specific needs, and potentially reduces the risk of introducing bias through subjective human interpretation. The ability to automatically generate rules from real-world data represents a significant step forward in making privacy-compliant medical data more readily available while maintaining its utility.
From Cancer Reports to Synthetic Rules: The Process Explained
The core of this new method involves extracting statistical patterns directly from existing cancer reports. Researchers begin by analyzing a corpus of de-identified patient records, specifically focusing on the prevalence and progression of various conditions described within those reports. This analysis goes beyond simply counting occurrences; it aims to capture relationships between factors like age, gender, family history, and the likelihood of specific diagnoses or treatments being administered. For example, they might determine that patients in a certain age range are significantly more likely to be diagnosed with a particular type of cancer, or that a specific treatment is commonly used for early-stage cancers.
This extracted statistical information isn’t directly usable by Synthea. Instead, it’s translated into what are called ‘Synthea rules.’ These rules provide instructions to the Synthea engine on how to generate synthetic patient data. A typical rule might state: ‘If a patient is between 60 and 70 years old, there is a 15% chance they will be diagnosed with breast cancer before age 75.’ The process involves mapping statistical findings from the cancer reports into this structured format. Crucially, this automation reduces the reliance on human experts who traditionally spend significant time crafting these rules based on their clinical knowledge.
Technically, this translation often utilizes probabilistic programming techniques and machine learning models to identify trends and generate rule suggestions. These suggestions are then validated against a smaller set of ‘gold standard’ rules created by medical professionals, ensuring the generated rules accurately reflect real-world patterns while maintaining data utility. The system iteratively refines the rule generation process based on this validation feedback, leading to increasingly precise and realistic synthetic patient data.
Glioblastoma as a Test Case & Future Implications
To illustrate the power of this automated rule generation process, researchers focused on glioblastoma, a particularly challenging cancer to model accurately. Glioblastoma’s complex progression, varying treatment responses, and dependence on numerous patient-specific factors make it an ideal test case for assessing the fidelity of synthetic data. The team meticulously validated the generated synthetic glioblastoma patient records against real-world datasets, focusing on critical aspects like disease course timelines – when tumors are detected, how quickly they progress, and typical survival rates. Statistical properties, such as age at diagnosis and response to standard treatments (like temozolomide), were also scrutinized to ensure the synthetic data mirrored observed patterns in the original patient populations.
The validation revealed that the synthetic glioblastoma data generally held up remarkably well against its real-world counterpart. While subtle discrepancies existed – for instance, slight variations in the distribution of certain biomarkers or minor differences in progression speeds across specific age groups – these were deemed acceptable given the inherent complexity of the disease and the limitations of any simplified modeling approach. Crucially, the synthetic data preserved key statistical trends and captured the overall heterogeneity seen in real glioblastoma patients, demonstrating that the automated rule generation effectively translated complex medical knowledge into a usable dataset.
Looking beyond glioblastoma, this methodology holds significant promise for generating synthetic patient data across a wide range of other diseases and conditions. Imagine creating synthetic datasets for rare genetic disorders, cardiovascular disease, or mental health conditions – areas where access to real-world data is often severely limited due to privacy concerns or logistical challenges. This could dramatically accelerate research into new treatments, diagnostic tools, and preventative strategies. Furthermore, the automated rule generation approach reduces the reliance on scarce expert knowledge, potentially democratizing synthetic data creation.
However, it’s important to acknowledge limitations. The accuracy of any synthetic dataset is inherently dependent on the quality and completeness of the underlying data used to inform the rules. Biases present in the original datasets will likely be reflected in the synthetic data, requiring careful mitigation strategies. Furthermore, while the approach captures statistical trends well, it may struggle to reproduce rare or highly idiosyncratic patient journeys that deviate significantly from established patterns. Ongoing research focuses on addressing these challenges and refining the automated rule generation process to produce even more realistic and useful synthetic patient data.
Validation: Does Synthetic Glioblastoma Data Hold Up?
To rigorously assess the utility of the generated synthetic glioblastoma patient data, researchers conducted a thorough validation process against the original dataset used to inform the rule creation. This involved comparing key aspects of disease progression between real and synthetic patients. Specifically, they analyzed survival curves, time-to-treatment metrics (like initial surgery or chemotherapy start), and the sequence of medical events – from diagnosis to follow-up scans. The goal was to ensure that the generated data mirrored the observed patterns in the original cohort, reflecting the complex temporal dynamics characteristic of glioblastoma.
Statistical properties were also scrutinized to confirm fidelity. Researchers compared distributions of demographic factors (age, gender), clinical variables (tumor size, KPS score – a measure of functional status), and laboratory values between the real and synthetic datasets. Overall, the validation showed strong agreement across these metrics; the synthetic data accurately replicated the statistical landscape of the original glioblastoma patient population. However, some minor discrepancies were noted in the precise representation of less frequent events or specific treatment pathways – areas where rule complexity may require further refinement to capture nuanced clinical realities.
Despite these minor deviations, the validation results strongly suggest that the automated approach successfully generates synthetic glioblastoma data with a high degree of representational accuracy. This demonstrates the potential for using such generated datasets not just for glioblastoma research but also for other complex diseases where access to real patient data is limited due to privacy concerns or scarcity.
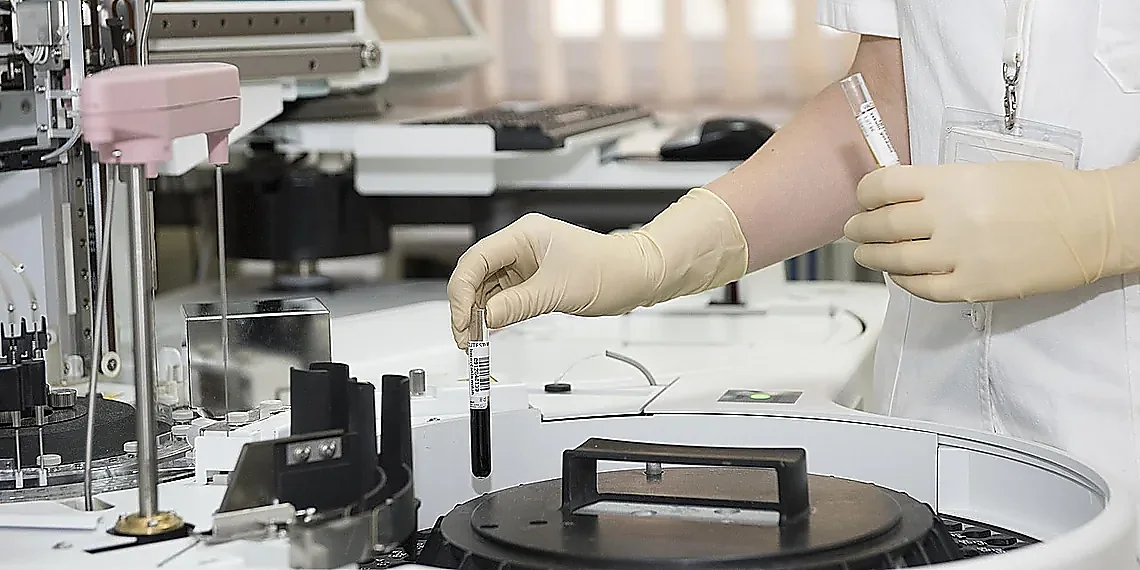

The convergence of AI and healthcare is rapidly transforming how we approach research, particularly concerning sensitive patient information. Automated generation techniques are no longer a futuristic concept; they’re delivering tangible benefits today, offering researchers unprecedented access to realistic datasets without compromising privacy. The potential for accelerating drug discovery, improving diagnostic accuracy, and personalizing treatment plans feels incredibly close thanks to advancements in this field. However, it’s crucial to remember that even the most sophisticated models produce data with inherent limitations; careful validation and critical assessment remain paramount when drawing conclusions from any dataset, including synthetic patient data. We must always acknowledge the assumptions baked into these algorithms and avoid over-generalizing findings. The future of healthcare innovation hinges on responsible adoption – embracing the power of AI while maintaining ethical standards and rigorous scientific scrutiny. To truly unlock this potential, a deeper understanding of the underlying methodologies is essential for all stakeholders, from researchers to policymakers. We strongly encourage you to delve further into tools like Synthea, which represent a significant step forward in generating realistic patient data. Explore how it could be applied within your own domain – whether you’re developing new algorithms, testing clinical workflows, or seeking to improve healthcare access and equity. Learn more about Synthea today and start envisioning the possibilities.
Your journey into the world of AI-driven healthcare research doesn’t have to end here; it’s just beginning!
Continue reading on ByteTrending:
Discover more tech insights on ByteTrending ByteTrending.
Discover more from ByteTrending
Subscribe to get the latest posts sent to your email.